- Department of Pharmaceutical Sciences, University of Toronto, 144 College Street, Toronto, ON, M5S 3 M2, Canada
- Imaging Research, Sunnybrook Research Institute, 2075 Bayview Avenue, Toronto, ON, M4N 3 M5, Canada
- Department of Medical Biophysics, University of Toronto, 610 University Avenue, Toronto, ON, M5G 2 M9, Canada
- STTARR Innovation Centre, Radiation Medicine Program, Princess Margaret Hospital, University Health Network, 101 College Street, Toronto, ON, M5G 1L7, Canada
- Department of Radiation Physics, Radiation Medicine Program, Princess Margaret Hospital, University Health Network, 610 University Avenue, Toronto, ON, M5G 2 M9, Canada
Received: 19 September 2009
Accepted: 3 February 2010
Published: 3 February 2010
© 2010 Cheung et al; licensee BioMed Central Ltd.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Few treatment options exist for patients with metastatic melanoma, resulting in poor prognosis. One standard treatment, dacarbazine (DTIC), shows low response rates ranging from 15 to 25 percent with an 8-month median survival time. The development of targeted therapeutics with novel mechanisms of action may improve patient outcome. Ribosome-inactivating proteins (RIPs) such as Shiga-like Toxin 1 (SLT-1) represent powerful scaffolds for developing selective anticancer agents. Here we report the discovery and properties of a single chain ribosome-inactivating protein (scRIP) derived from the cytotoxic A subunit of SLT-1 (SLT-1A), harboring the 7-amino acid peptide insertion IYSNKLM (termed SLT-1AIYSNKLM) allowing the toxin variant to selectively target and kill human melanoma cells.
SLT-1AIYSNKLM was able to kill 7 of 8 human melanoma cell lines. This scRIP binds to 518-A2 human melanoma cells with a dissociation constant of 18 nM, resulting in the blockage of protein synthesis and apoptosis in such cells. Biodistribution and imaging studies of radiolabeled SLT-1AIYSNKLM administered intravenously into SCID mice bearing a human melanoma xenograft indicate that SLT-1AIYSNKLM readily accumulates at the tumor site as opposed to non-target tissues. Furthermore, the co-administration of SLT-1AIYSNKLM with DTIC resulted in tumor regression and greatly increased survival in this mouse xenograft model in comparison to DTIC or SLT-1AIYSNKLM treatment alone (115 day median survival versus 46 and 47 days respectively; P values < 0.001). SLT-1AIYSNKLM is stable in serum and its intravenous administration resulted in modest immune responses following repeated injections in CD1 mice.
These results demonstrate that the evolution of a scRIP template can lead to the discovery of novel cancer cell-targeted compounds and in the case of SLT-1AIYSNKLM can specifically kill human melanoma cells in vitro and in vivo.
The incidence of melanoma has been rising in the United States for the past sixty years(1)(2). Despite prevention efforts, it remains the second leading cause of lost productive years among all cancers, and is responsible for more than 7,000 deaths annually(2). Novel melanoma-targeted therapeutic agents are needed to improve prognosis, since traditional treatments such as dacarbazine (DTIC) and IL-2 only yield a 5% survival advantage of more than five years for patients with advanced melanoma(2). Currently, targeted agents such as monoclonal antibodies and recombinant proteins account for more than a quarter of all cancer therapeutics that have been newly approved or are presently in clinical trials(3)(4). Although effective in delaying the progression of certain cancers, naked antibodies such as Herceptin and Avastin do not cure the disease(5)(6)(7). This limitation has led to the use of tumor-targeted protein ligands in delivering potent therapeutic cargoes such as radionuclides, drugs, and protein toxins to tumor sites, bringing the overall concept of personalized medicine closer to reality(3)(4). In particular, immunotoxins and protein toxin conjugates have been the subject of clinical trials with a fusion construct of IL-2 to diphtheria toxin (Ontak) being approved by the FDA for the treatment of cutaneous T-cell lymphoma. Nevertheless, the use of toxin conjugates as successful cancer therapeutics remains limited(8)(9)(10)(11)(12). To address some of the design challenges facing targeted protein toxins, we created a combinatorial protein library based on the cytotoxic domain of a protein toxin with a view to directly screen in cell-based assays for variants bearing new ligand specificities and able to selectively destroy cancer cells(13). The approach makes use of the cytotoxic A subunit of a bacterial ribosome-inactivating protein (RIP), namely Shiga-like Toxin 1 (SLT-1), as a protein scaffold to design anticancer agents. Specifically, the SLT-1 A subunit encodes all functions necessary to route itself out of cellular organelles in order to reach and inactivate ribosomes present in the cytoplasm of eukaryotic cells. This event subsequently leads to apoptosis(14). However, the SLT-1 A subunit lacks the ability to target cancer cells. In this report, we have inserted a random 7-amino acid peptide motif into the structure of the SLT-1A domain to create a combinatorial library of this protein template expressing toxic SLT-1A mutants harboring a putative peptide ligand that may specifically recognize, enter and kill cancer cells (Figure 1). Searching through such a library yielded a SLT-1A variant termed SLT-1AIYSNKLM that selectively targets human melanoma cell lines. The mechanism of action and therapeutic activity of this single chain ribosome-inactivating protein (scRIP) variant closely parallel the predicted properties of a targeted ribosome-inactivating protein, suggesting that RIP A subunit libraries may represent a useful discovery tool for targeted protein-based therapeutics. We have termed this discovery approach RESCRIPT (Rapid Evolution and Selection of Cancer-specific Ribosome-Inactivating Protein Toxins) (see Additional File 1: Figure S1) which is based on the identification of targeted RIPs with no a priori knowledge of surface markers associated with cancer cells.
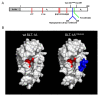
(A) Schematic diagram of the SLT-1 A subunit. Furin cleavage occurs within its protease-sensitive loop (C242 to C261) between residues 251 and 252 (FCS 251/252) of the A subunit(44). A random 7-amino acid peptide library was inserted between residues 245 and 246, generating cytotoxic A subunit variants with a surface-displayed library insert. Catalytic residues are shown in red. (B) Modeled surface representations of the SLT-1 A subunit with and without a 7-residue long insertion. The models were derived from the crystal structure of Shiga Toxin(45). The peptide insertion (blue) is solvent-exposed and positioned away from residues involved in catalysis (residues Y77, Y114, E167, R170, W203 colored in red)(20)(46)(47). Models were rendered using PyMOL Software Version 1.0r1.
Click image to view larger.
Format: TIFF Size: 4MB Download file
Diagram outlining the general procedure for the screening and identification of Shiga-like Toxin 1 (SLT-1) A subunit toxin variants with novel cancer-targeting and killing properties. His-tagged toxin variants were individually purified and applied to 96-well plates seeded with cancer cell lines. Cell viability was assessed using a sulforhodamine B (SRB) assay(24).
Format: DOC Size: 71KB Download file
This file can be viewed with: Microsoft Word Viewer
A combinatorial SLT-1A protein library was constructed by inserting a random 7-amino acid peptide element between residues 245 and 246 of the A subunit (Figure 1A and Additional File 2). The insertion of this heptapeptide did not affect the catalytic activity of SLT-1A (Additional File 3). This peptide element was also shown to be exposed on the surface of the A subunit (Additional File 3). Single bacterial colonies were picked from the library and led to the purification of 9,400 His-tagged scRIP variants that were tested for their ability to kill the wt SLT-1-resistant human melanoma cell line 518-A2 (see Additional File 1: Figure S1 and Additional File 2). The initial screens led to the recovery of 112 SLT-1A toxin variants displaying cytotoxic profiles towards cancer cell lines. These SLT-1A variants were re-screened against 518-A2 cells, as well as against a panel of twelve other cell lines (human unless otherwise indicated): PC-3 (prostate cancer), SKBR-3 (breast cancer), CAMA-1 (breast cancer), U87 (glioma), OVCAR-3 (ovarian carcinoma), SiHa (cervical cancer), PanC (pancreatic cancer), B16-F10 (mouse melanoma), Vero (monkey, normal kidney), HS-216 (normal fibroblast), H-MEC (normal mammary epithelial cells), and H-REC (normal kidney cells). This counter-selection step led to the identification of an A subunit toxin variant, named SLT-1AIYSNKLM, harboring the peptide sequence IYSNKLM, that selectively targets and kills 518-A2 cells in a dose-dependent manner [CD50 ~300 nM] (Figure 2A). Furthermore, SLT-1AIYSNKLM demonstrates broad specificity towards melanoma cell lines, killing 7 of 8 human melanoma cell lines tested (518-A2, A-2058, A-375, C-32, MALME-3 M, MeWo, SK-Mel-2, and SK-Mel-28), as well as human melanocytes to a lesser extent (Figure 2B).
Format: TIFF Size: 5.5MB Download file
(A) The heptapeptide insert did not affect the ribosome-inactivating activity of the purified toxin variants compared to the wt A subunit as measured by the biosynthesis of luciferase (relative light units) in a coupled transcription/translation assay. Legend: wt SLT-1 (open squares), SLT-1APDTRPAP (open triangles), and a catalytically-inactive SLT-1 variant bearing E167A and R170A mutations within the A subunit (open circles). The peptide insert within the context of the toxin A subunit is exposed and recognized by large proteins such as antibodies. Samples were probed with either an anti-SLT-1 A subunit polyclonal antisera or a PDTRPAP-specific anti-Onc-M27 monoclonal antibody. (B) Western blot analyses of purified A and A1 subunits for wt SLT-1 or SLT-1APDTRPAP. (C) ELISA experiments comparing the immunoreactivity of purified AB5, A and A1 subunits to the Onc-M27 mAb for wt SLT-1 and SLT-1APDTRPAP toxins.
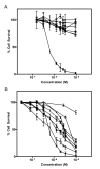
(A) Dose-response curves illustrating the specific cytotoxicity of SLT-1AIYSNKLM (black diamonds, black triangles, black squares, black circles) compared to wt SLT-1A (white diamonds, white triangles, white squares, white circles) for the 518-A2 human melanoma cell line (black squares, white circles). The carcinoma cell lines CAMA-1 (black diamonds, white diamonds; human breast), HepG2 (black triangles, white triangles; human liver), and PC-3 (black squares, white squares; human prostate) are insensitive to SLT-1AIYSNKLM. (B) Dose-response curves illustrating the cytotoxicity profile of SLT-1AIYSNKLM towards 8 different human melanoma cell lines: 518-A2 (black circles), A-375 (black inverted triangles), SK-Mel-28 (white circles), MeWo (black squares), A-2058 (white inverted triangles), MALME-3 M (white triangles), SK-Mel-2 (white squares), and C-32 (black triangles) as well as human melanocytes (white diamonds). Error bars represent s.e.m. for experiments performed in quadruplicate.
Click image to view larger.
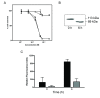
(A) Dose-response curves indicating a comparable cytotoxicity for the purified, furin-cleaved A subunit of SLT-1AIYSNKLM (A1 and A2 subunits remain linked by a disulfide bridge between C242 and C261; white squares) and the A1 subunit alone of SLT-1AIYSNKLM (white circles) towards 518-A2 cells. The catalytically-inactive forms (Y77S) of these same molecules (black squares, black circles) were not cytotoxic towards 518-A2 human melanoma cells at concentrations of up to 10 μM. (B) Western blot analysis demonstrating the cleavage of PARP following treatment of 518-A2 cells with SLT-1AIYSNKLM. (C) Effect of exposing 518-A2 (black bars) and PC-3 cells (grey bars) to SLT-1AIYSNKLM on the activation of caspase-3 as measured using a fluorescent peptide substrate for caspase-3. PC-3 cells are insensitive to SLT-1AIYSNKLM. Error bars represent s.e.m. for experiments performed in quadruplicate.
Click image to view larger.
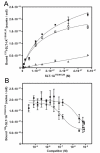
(A) Specific binding (white squares), total binding (black squares), and non-specific binding (white triangles) of 125I-SLT-1AIYSNKLM to 518-A2 cells at 4°C. (B) Displacement curves in which 518-A2 cells were treated with 45 nM of 125I-SLT-1AIYSNKLM in the presence of increasing concentrations of either unlabeled SLT-1AIYSNKLM (black circles) or a synthetic 16-residue peptide containing the inserted 7-amino acid peptide ligand (HHHIYSNKLMASRVAR) (black squares). Data points represent the s.e.m. of experiments performed in triplicate.
Click image to view larger.
Format: TIFF Size: 5.5MB Download file
Western blot analysis demonstrating the serum stability of SLT-1AIYSNKLM at 37°C over a period of 24 h.
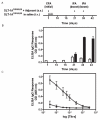
(A) Injection schedule of SLT-1AIYSNKLM with and without adjuvant. Treatments consisted of i.v. injections of SLT-1AIYSNKLM prepared in saline into CD1 mice as a series of 5 consecutive (daily) tail vein injections or a single s.c. injection of SLT-1AIYSNKLM in Complete Freund's adjuvant (CFA) followed by two s.c. boosts of the antigen in Incomplete Freund's adjuvant (IFA) at days 21 and 35. (B) Histogram illustrating the average IgG immune responses (1:1000 titer dilution) from 3 CD1 mice injected with SLT-1AIYSNKLM in the presence (black bars) or absence of adjuvant (white bars) as measured by ELISA at 405 nm. (C) Representative IgG immune responses (average ELISA signals at 405 nm versus antisera titers; day 42) engendered for groups of three CD1 mice after injection with SLT-1AIYSNKLM in the presence (white circles) or absence of adjuvant (black circles). Data represent the s.e.m. of experiments performed in triplicate.
Click image to view larger.
125I-SLT-1AIYSNKLM was subsequently injected i.v. into SCID mice bearing 518-A2 tumor xenografts(23) to establish the pharmacokinetic parameters of this agent and its ability to localize into tumor cells. The scRIP was rapidly cleared through the kidneys with a circulating half-life (t1/2) of ~11 min, a value that is consistent with a protein of this size (Figure 6A)(24). Importantly, the biodistribution profile showed marked tumor uptake and retention of 125I-SLT-1AIYSNKLM, reaching a maximum of ~37% injected dose per gram of tumor at 1 h post-i.v. injection (Figure 6B). Tissue analyses also demonstrated the selectivity of 125I-SLT-1AIYSNKLM in that tumor localization increases as the molecule is eliminated more slowly from the tumor than from blood and other tissues. Specifically, the tumor-to-blood ratios of 125I-SLT-1AIYSNKLM at 1 h, 6 h, 12 h, and 24 h post-injection are 6.6, 3.1, 5.9 and 3.0 respectively.
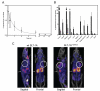
(A) Blood clearance of 125I-SLT-1AIYSNKLM shown as percentage of injected dose per mL blood collected at various time points over a 72 h period post-i.v. injection. The clearance rate (t1/2) of 125I-SLT-1AIYSNKLM was shown to be ~11 min. (B) Biodistribution per collected gram of wet tissue of 125I-SLT-1AIYSNKLM after 1 h (black bars), 6 h (light grey bars), 12 h (dark grey bars) and 24 h (white bars) post-i.v. administration. (C) Tumor localization of 125I-SLT-1AIYSNKLM versus wt 125I-SLT-1A (negative control) as shown by nanoSPECT and CT imaging. The composite images show the tissue uptake of the radiolabeled proteins (as pink colored areas) as well as the location of tumor xenografts (white circles) and that of kidneys (grey arrows).
Click image to view larger.
To visualize the tumor-localization of SLT-1AIYSNKLM, a group of 518-A2 xenograft-bearing SCID mice received a single i.v. injection of either 125I-SLT-1AIYSNKLM (900 μCi) or wt 125I-SLT-1A (900 μCi; negative control). The distribution of radiolabeled scRIPs was recorded 45 min post-injection with a nanoSPECT/CT scanner. The resulting CT MIP (Maximum Intensity Projection) images were then overlaid with the SPECT image slice that transversed the central plane of the tumor (Figure 6C). These images highlight the rapid localization and specificity of 125I-SLT-1AIYSNKLM into 518-A2 cell xenografts as compared to the control, wt 125I-SLT-1A, which shows no tumor localization.
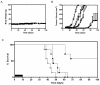
k circles), or a 5-day course of DTIC (i.p. 8 mg/kg) (white squares), or the combination of both SLT-1AIYSNKLM (i.v. 0.5 mg/kg) with DTIC (i.p. 8 mg/kg) regimens (black squares), (n = 7). (A) Mice showed no change in body weight related to treatment regimens over time. (B) Measured tumor volumes of mice demonstrate a significant synergistic effect of combining SLT-1AIYSNKLM and DTIC treatments when compared to either treatment alone or to the saline control (P < 0.0001). (C) Kaplan-Meier plot comparing animal survival as a function of treatment regimens (ten-day regimen; black bar).
Click image to view larger.
The incidence of melanoma has been rapidly rising worldwide with no effective treatment in sight for patients with metastatic forms of this disease(1). Despite decades of evaluating new therapeutic modalities for advanced melanoma, patients are mainly being treated with agents that display low response rates, i.e. DTIC and IL-2(2). In view of the urgent need for new therapeutic strategies, we designed and screened a combinatorial library based on the scaffold of a single chain ribosome-inactivating protein (scRIP), namely the cytotoxic A subunit of SLT-1, to identify SLT-1 A variants that target and kill human melanoma cells. In this study, we demonstrated that the screening of this combinatorial SLT-1A protein library led to the identification of a toxin variant termed SLT-1AIYSNKLM that selectively binds to human melanoma cells, is internalized by them and results in their apoptotic death.
Immunotoxins derived from the fusion of the cytotoxic domain of Pseudomonas exotoxin A (PE) to a tumor-targeted Fv fragment, have been shown to kill cells using a combination of cytotoxic and cytostatic mechanisms of action, namely the induction of apoptosis and cell death (cytotoxic mechanism) due to the inhibition of protein synthesis (cytostatic effect)(25)(26). Wild type SLT-1 utilizes both mechanisms. Firstly, the SLT-1 A subunit undergoes retrograde transport to the endoplasmic reticulum, where it then retrotranslocates to the cytoplasm and halts protein synthesis via its catalytic activity by cleaving 28S rRNA(27). The blockage of protein synthesis subsequently triggers apoptosis through the activation of caspases 8, 9, and 3 leading to the display of morphological characteristics such as membrane blebbing, DNA fragmentation, chromatin condensation and cell shrinkage(14)(17). As presented in Figure 3, only the catalytically active SLT-1AIYSNKLM kills cells, activates caspases-3 and cleaves PARP, as observed for ricin, another common RIP family member(28). Thus, one key advantage of SLT-1A as a combinatorial library template is that it yields small scRIP agents with short new ligand binding domains without altering the inherent intracellular localization and cytotoxic properties of the A subunit. This mechanism of action is distinct from most commonly prescribed anticancer agents in the clinic, suggesting that the cytotoxic A subunit of SLT-1 could complement the action of existing drugs in combination therapy.
The melanoma-specific scRIP SLT-1AIYSNKLM was directly derived using RESCRIPT (Additional File 1: Figure S1) from a cell-based screen and its therapeutic potential confirmed in vivo without the need to re-engineer parts of its structure. This discovery and optimization approach differs from preexisting immunotoxin design strategies. Specifically, the traditional two-step assembly of immunotoxins and related conjugates has been based on an a priori selection of a known tumor marker. However, the fusion or chemical coupling of a protein ligand (typically > 25 kDa) to a toxin domain often generates large protein constructs displaying altered functions in terms of cell targeting, cellular processing and cytotoxic functions(9)(22)(29). Secondly, RESCRIPT, as a discovery tool, is distinct from phage display approaches which are typically used to identify peptide and protein ligands and are not presently compatible with cell killing assays. RESCRIPT thus limits the need for post-discovery, protein engineering steps required to ensure that functions such as cell targeting, cellular routing and toxicity are retained by the resulting conjugates.
Mechanistically, SLT-1AIYSNKLM behaves as a catalytically active RIP causing apoptosis in 518-A2 melanoma cells (Figure 3). SLT-1AIYSNKLM was shown to bind to a surface receptor on 518-A2 cells with a Kd of 18 nM. These cells express about 140,000 copies of the receptor. More importantly, the specific binding of this A subunit variant to receptors on melanoma cells was rapid, saturable and reversible (Figure 4). Preliminary biochemical studies aimed at determining the nature of the surface determinant recognized by SLT-1AIYSNKLM on 518-A2 melanoma cells, suggests that the receptor may not be a protein. Specifically, pre-treating 518-A2 cells extensively with trypsin yielded an identical SLT-1AIYSNKLM cell cytotoxicity profile to that of untreated 518-A2 cells. Furthermore, preliminary cell surface radioiodination followed by crosslinking/pull-down experiments with His-tagged SLT-1AIYSNKLM and wt SLT-1A did not reveal any unique radiolabeled membrane species that may act as putative SLT-1AIYSNKLM receptors on SDS-PAGE. Both approaches have limitations in terms of specificity and detection sensitivity. Biochemical and proteomic approaches are on-going to define this melanoma-specific receptor. Studies are also being conducted to determine whether the SLT-1AIYSNKLM receptor may be a cell-surface melanoma marker that has already been described in the literature.
Although several melanoma biomarkers have previously been reported, only a few are expressed on the cell-surface and therefore would qualify as possible SLT-1AIYSNKLM receptor candidates(30). These surface markers include HMW-MAA (high molecular weight melanoma associated antigen; melanoma chondroitin sulfate proteoglycan; MCSP), S100B (originally known as S100), CD44, CXCR4 (CXC chemokine receptor) and CEACAM1 (carcinoembryonic-antigen-related cell adhesion molecule 1)(31)(32)(33)(34)(35)(36)(37). However, these known markers are present on normal tissues as well as other cancer types, highlighting challenges in designing melanoma-specific therapies through the traditional method of conjugating a ligand to a cell-killing component (30)(33)(36)(38)(39). Nevertheless, antibodies raised against some of these biomarkers, such as anti-Id mAb MK2-23 for HMW-MAA, have shown promise in increasing patient survival. To date, there is still a lack of clinically successful melanoma-specific markers, and none for the detection of primary tumors for high risk patients(30)(40)(41).
The pharmacological properties of SLT-1AIYSNKLM were also investigated in a mouse model and suggest that the molecule is cleared with the expected half-life profile of antibody fragments with a mass of ~30 kDa. SLT-1AIYSNKLM is also stable in human serum, displays modest immunogenicity, and accumulates readily in tumor xenografts of 518-A2 cells implanted in SCID mice (Figure 6, Figure S4 and Figure 5 and 7). This scRIP also retains its cytotoxic activity both in vitro (Figure 2 and 3) and in vivo as shown by the level of tumor regression observed upon treating tumor-bearing mice (Figure 7). These remarkable properties suggest that SLT-1AIYSNKLM be pursued as a possible treatment modality for melanoma patients displaying metastatic forms of this disease.
We have developed and screened a combinatorial single chain, ribosome-inactivating protein (scRIP) library with a view to use the intrinsic cytotoxicity of all members of the library (built-in search engine) as a tool to identify melanoma-specific scRIP variants in the context of cell viability assays. Library searches led to the discovery of SLT-1AIYSNKLM, a novel scRIP variant of the cytotoxic A subunit of Shiga-like Toxin 1 that is able to target and specifically kill human melanoma cells. SLT-1AIYSNKLM represents a candidate agent for treating patients with metastatic melanoma.
Format: TIFF Size: 54KB Download file
Coomassie-stained, SDS-PAGE gel showing Lane 1) purified SLT-1AIYSNKLM before furin digestion; Lane 2) purified SLT-1AIYSNKLM after furin treatment; 3) purified wt SLT-1A before furin digestion; Lane 4) purified wt SLT-1A after furin treatment.
Immunogenicity studies were performed with CD1 mice (3 mice per time point) given a series of 5 consecutive (250 μg/kg; 100 μL daily) i.v. injections of SLT-1AIYSNKLM prepared in saline to mimic the therapeutic treatment procedure. As a positive control, a single dose (250 μg/kg; 100 μL) of this A chain emulsified (1:1) in Complete Freund's adjuvant (CFA) was injected subcutaneously into CD1 mice (control group). The control animals also received s.c. injections of the same antigen emulsified (1:1) in Incomplete Freund's adjuvant (IFA; 250 μg/kg; 100 μL dose) at day 21 and 35. To assess IgG immune responses raised against SLT-1AIYSNKLM, blood samples were collected at various time intervals post-injection (day 1, 10, 15, 21, 35, 42 for SLT-1AIYSNKLM; day 21, 35, 42 for the antigen with adjuvant). IgG responses were titrated for each mouse by serially diluting serum samples and measuring IgG levels by ELISA in 96-well plates pre-coated with 100 ng of SLT-1AIYSNKLM. Serum-treated wells (in triplicate) were subsequently exposed to an HRP-conjugated goat-anti-mouse IgG (1:1000) and ELISA signals recorded with a plate reader at 405 nm using the substrate ABTS (2,2'-Azinobis [3-ethylbenzothiazoline-6-sulfonic acid]-diammonium salt).
Imaging experiments were carried out at the UHN STTARR facility using a dual-modality NanoSPECT/CT (Bioscan) and are further described under Additional File 2 (2 mice). All animal protocols were reviewed and approved by the Animal Care Committee at the University Health Network.
(RIP): Ribosome-Inactivating Protein; (scRIP): Single Chain Ribosome-Inactivating Protein; (SLT-1): Shiga-Like Toxin 1; (SLT-1A): Shiga-Like Toxin 1 A subunit; (SLT-1AIYSNKLM): Shiga-Like Toxin 1 A subunit with an IYSNKLM insertion between residues 245 and 246; (DTIC): Dacarbazine; (PARP): Poly(ADP) Ribose Polymerase; (MIP): Maximum Intensity Projection; (CBCT): Cone-Beam CT; (OSEM): Ordered Subset Expectation Maximization; (PE): Pseudomonas exotoxin A; (HMW-MAA): High Molecular Weight Melanoma Associated Antigen; (MCSP): Melanoma Chondroitin Sulfate Proteoglycan; (CXCR4): CXC chemokine receptor; (CEACAM1): Carcinoembryonic-Antigen-Related Cell Adhesion Molecule 1
MCC, SP, XW, DEG and AAW declare that they have no competing interests. RK formerly received a salary from Molecular Templates Inc. LR and JG are founding members of Molecular Templates Inc. Patent applications have been filed for SLT-1AIYSNKLM by the University Health Network (UHN, Toronto, academic entity of JG). UHN recently sold these patent rights to Molecular Templates Inc.
JG conceived the study, participated in its design and coordination, and helped draft the manuscript. MCC designed, carried out and performed the relevant data analysis for the protein purification, cell binding, pharmacokinetics, biodistribution, imaging, densitometry, and serum stability experiments, conducted part of the mouse survival studies, generated the summary figures for the library design and screening, graphed and analyzed the data for the cell cytotoxicity, immunogenicity, and mouse survival studies, and helped draft the manuscript. LR participated in the general supervision of the research group while they collected data for the cell cytotoxicity, immunogenicity, and mouse survival studies. SP designed the preliminary tripeptide library and developed screening approaches, carried out apoptosis experiments as well as performed the relevant data analysis. XW designed and carried out the heptapeptide library design and screening, as well as the initial cell survival studies and identification of the tumour-targeted SLT-1A variant. RK carried out and collected data for the mouse survival and immunogenicity studies as well as performed the protein purification and heptapeptide library screening. DEG participated in the design, coordination, execution, and data analysis for the mouse imaging studies. AAW helped coin the acronym RESCRIPT and assisted in data collection for some of the mouse biodistribution work. All authors read and approved the final manuscript.
This study was supported by the Canadian Cancer Society Research Institute. We also would like to thank Dr. Gregory M.K. Poon for assisting with the radiolabeling, cell-binding experimental design and data analysis, Eric Huang for the optimization of protein production and purification, and for helping with pharmacokinetic, biodistribution, imaging, and mouse survival studies, Erin Green, and Qing Hong Zhang, for the production and purification of proteins and assistance with mouse survival and immunogenicity work, Amaalia E. Broad and Andrew J. McCluskey for their technical expertise and help with mouse work, Doug Vines for assistance and advice with respect to mouse imaging, and Dr. Nenad Cicmil for help with protein modeling software. We also thank Dr. Raymond Reilly and members of his laboratory who engaged in helpful discussions with respect to the design and data analysis relating to our radiolabeling studies.


